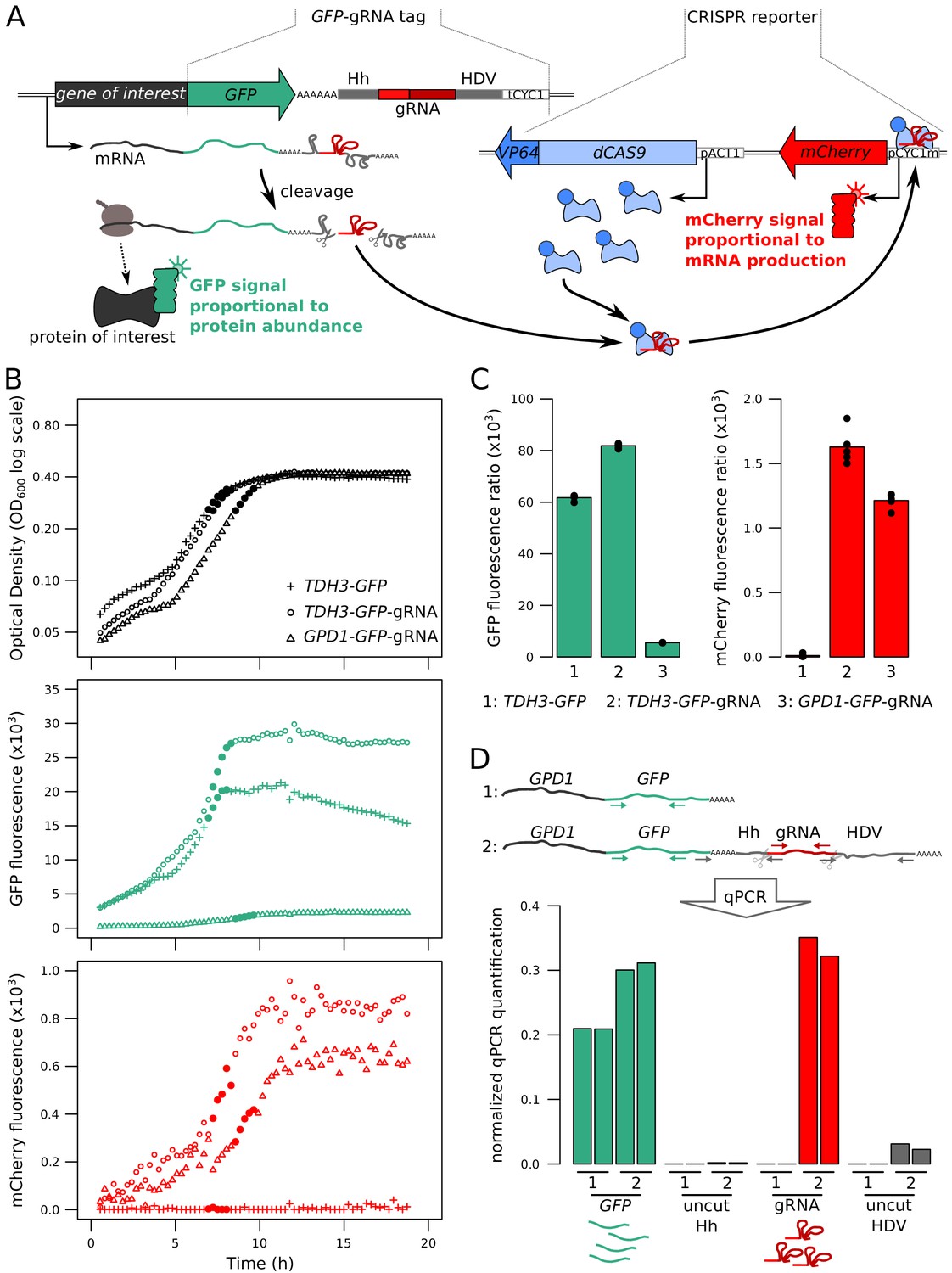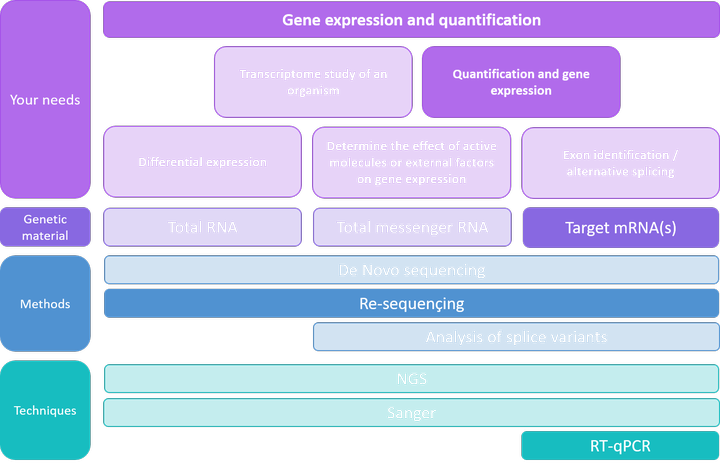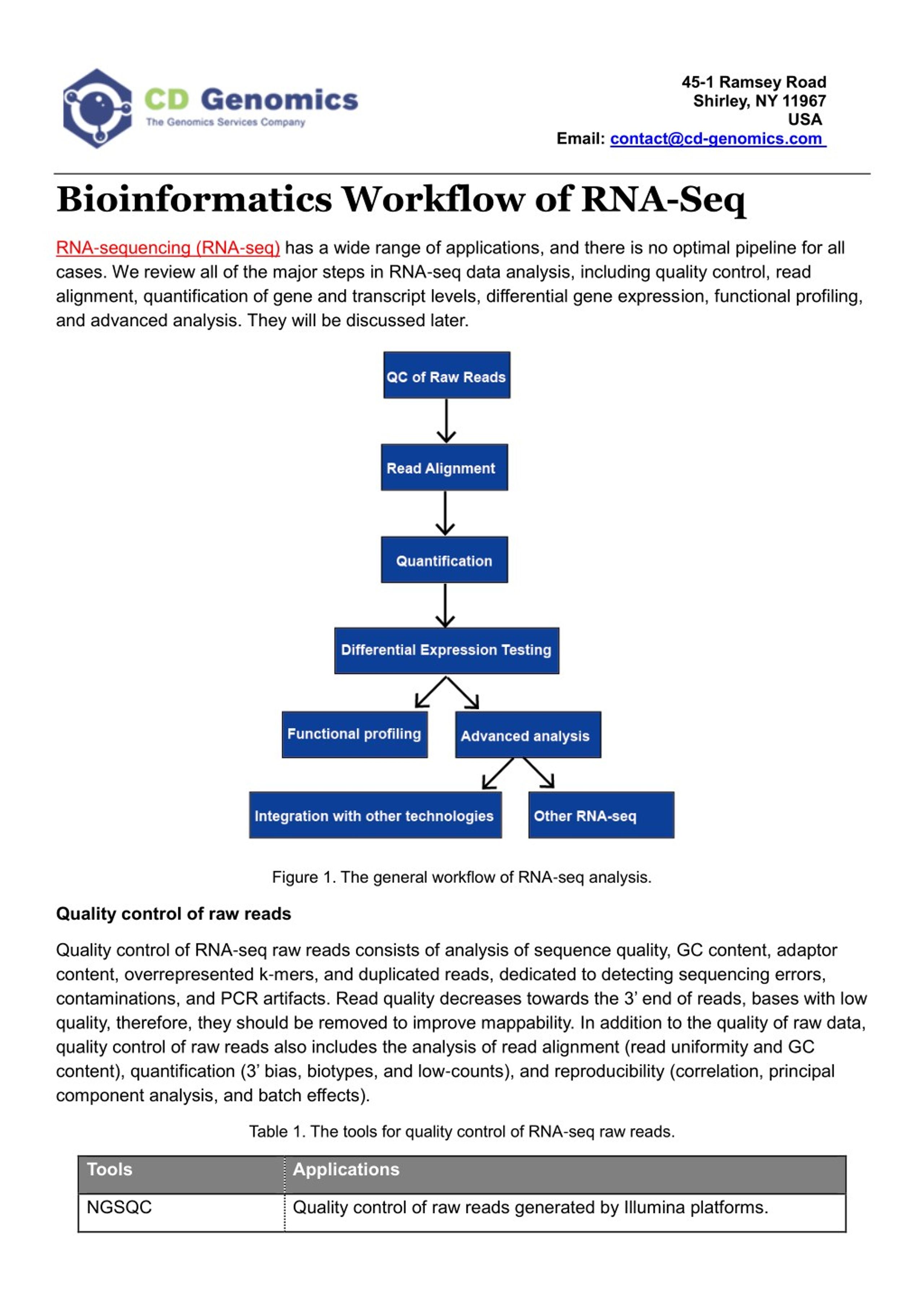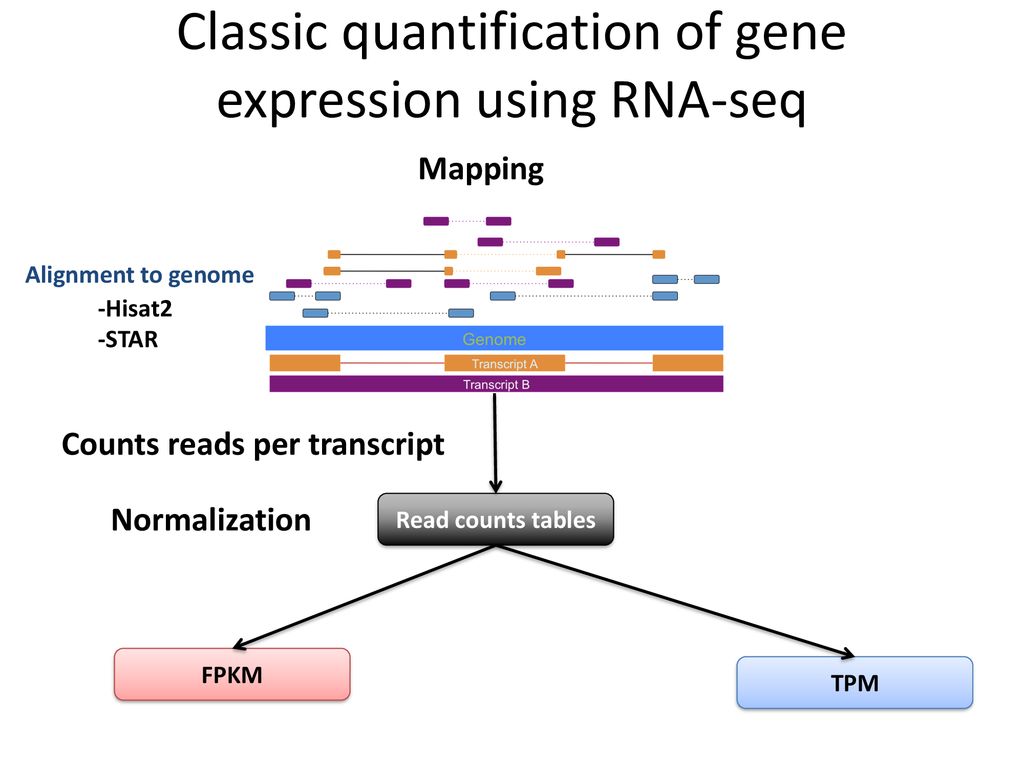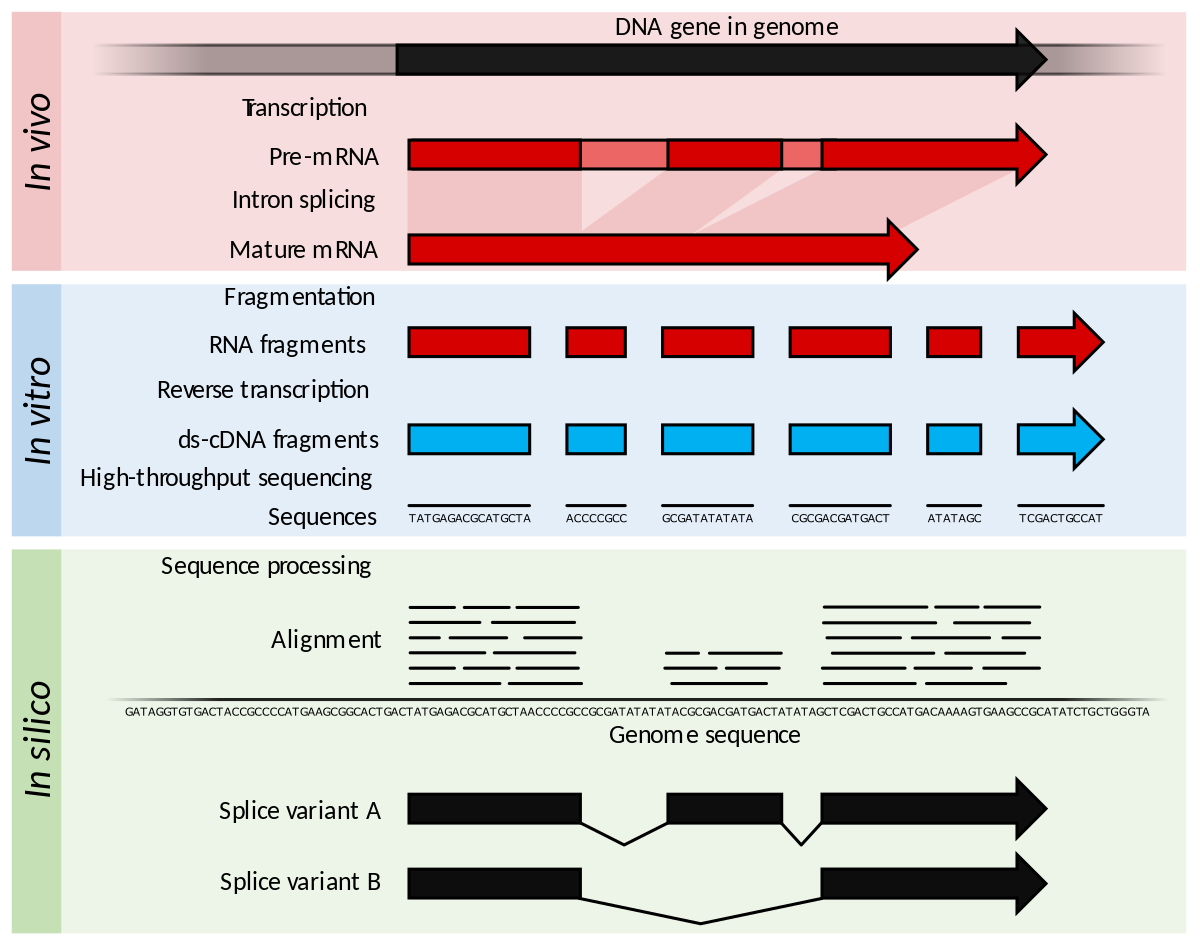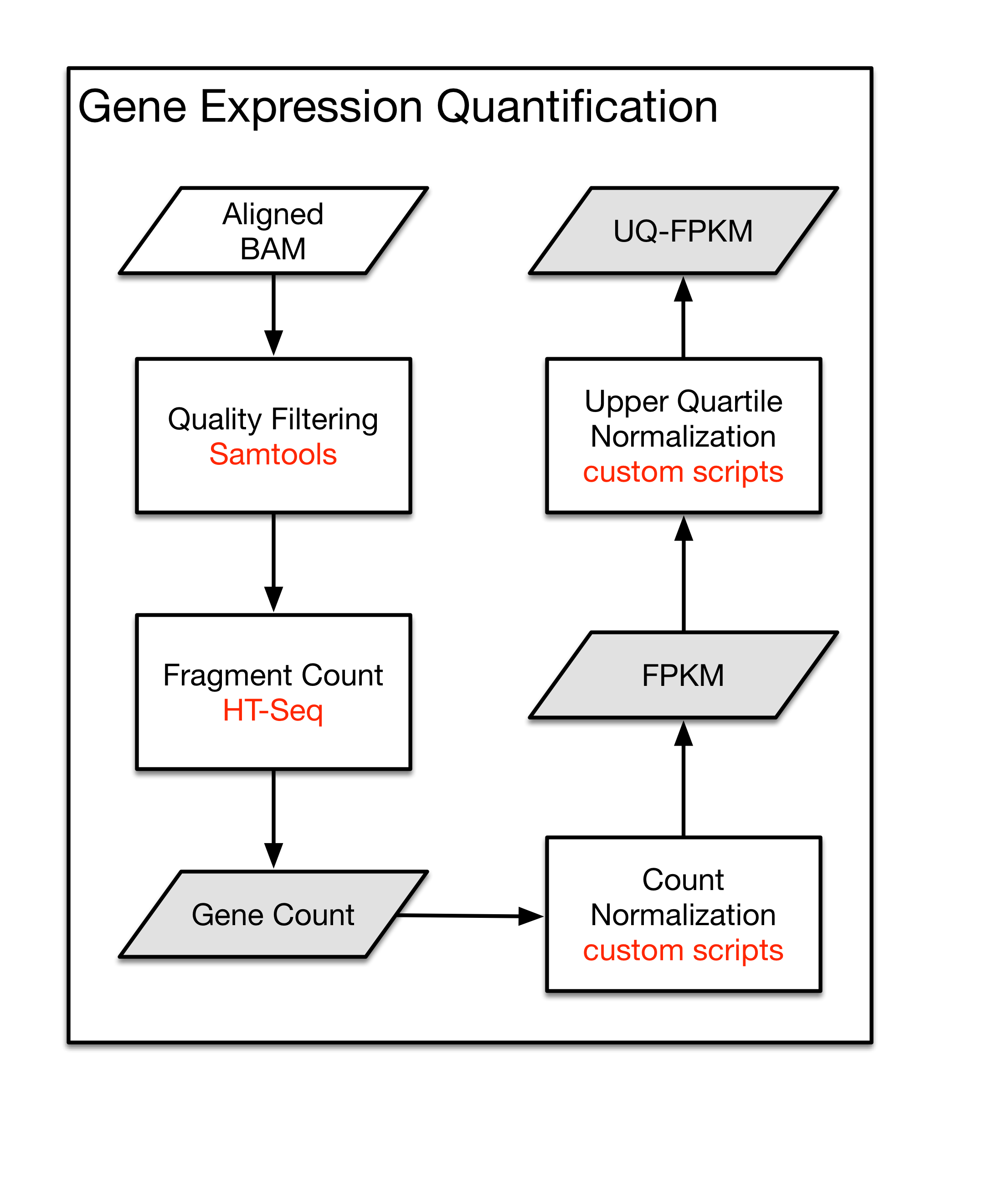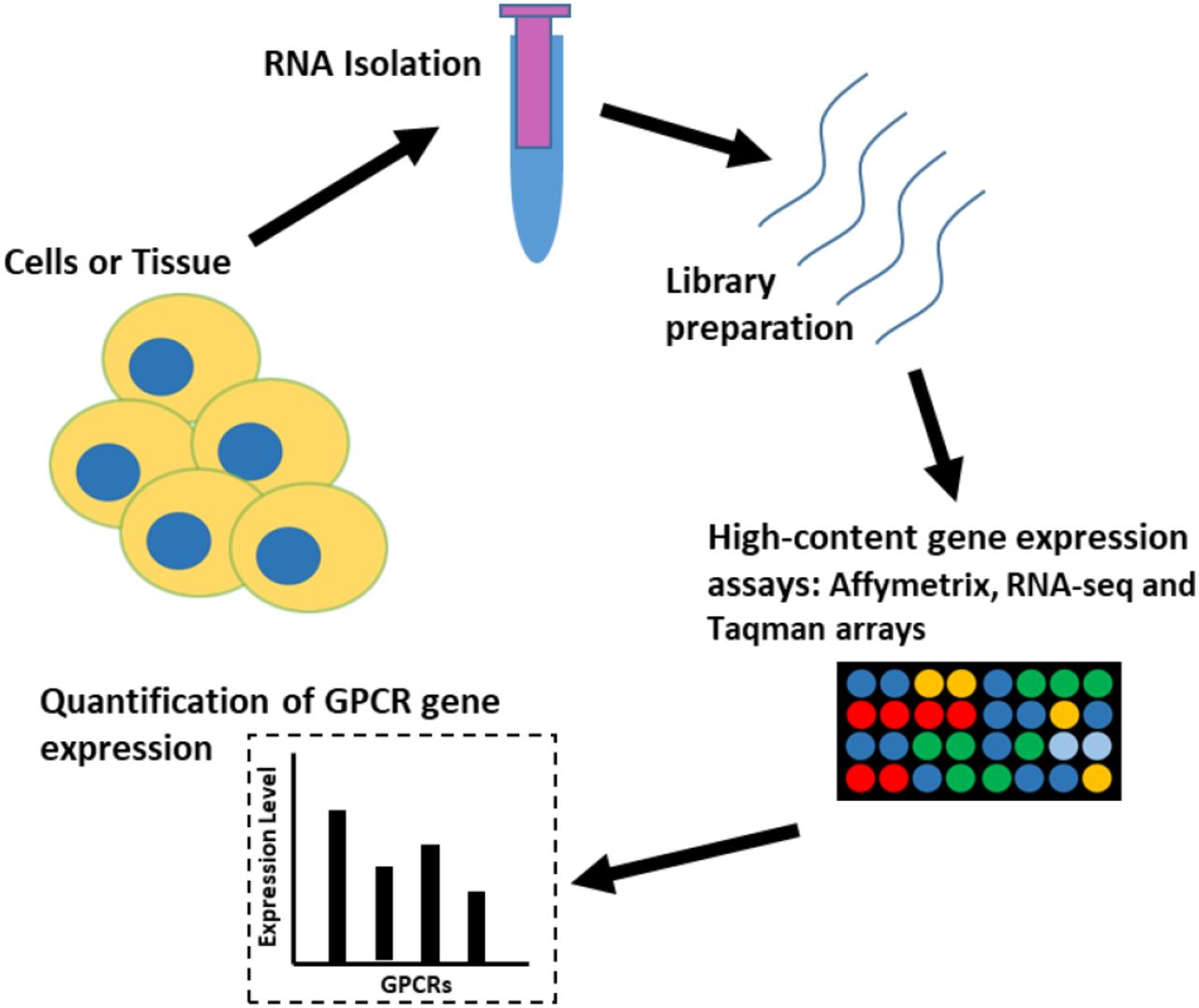
Detection and quantification of GPCR mRNA: An assessment and implications of data from high-content methods | bioRxiv

Standardization of Gene Expression Quantification by Absolute Real-Time qRT-PCR System Using a Single Standard for Marker and Reference Genes - Yi-Hong Zhou, Vinay R. Raj, Eric Siegel, Liping Yu, 2010
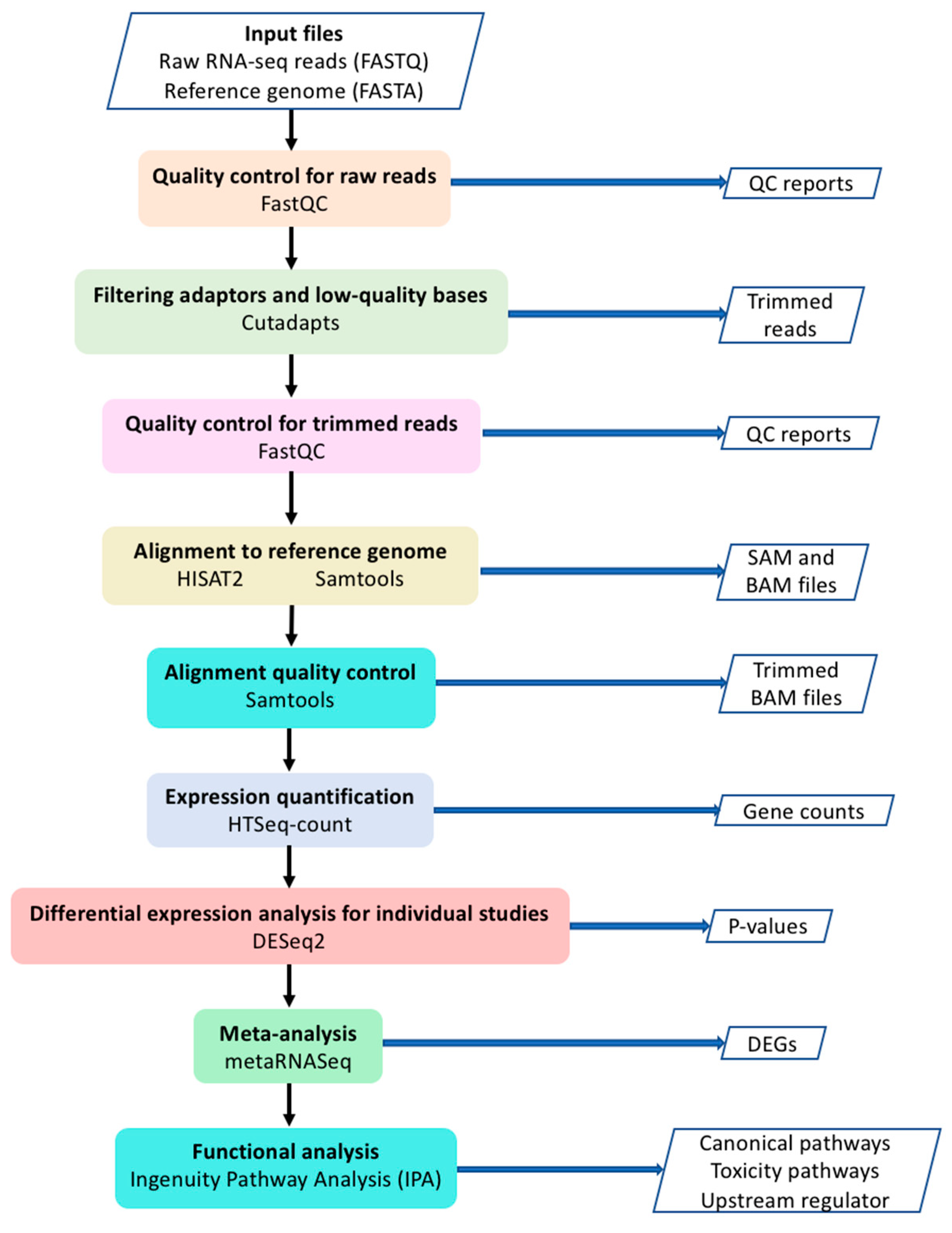
Genes | Free Full-Text | Meta-Analysis of Dilated Cardiomyopathy Using Cardiac RNA-Seq Transcriptomic Datasets

RNA-Seq Quantification of Hepatic Drug Processing Genes in Germ-Free Mice | Drug Metabolism & Disposition
![Validation of reference genes for gene expression studies in tartary buckwheat (Fagopyrum tataricum Gaertn.) using quantitative real-time PCR [PeerJ] Validation of reference genes for gene expression studies in tartary buckwheat (Fagopyrum tataricum Gaertn.) using quantitative real-time PCR [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/6522/1/fig-3-full.png)
Validation of reference genes for gene expression studies in tartary buckwheat (Fagopyrum tataricum Gaertn.) using quantitative real-time PCR [PeerJ]

Quantification of differential gene expression by multiplexed targeted resequencing of cDNA | Nature Communications
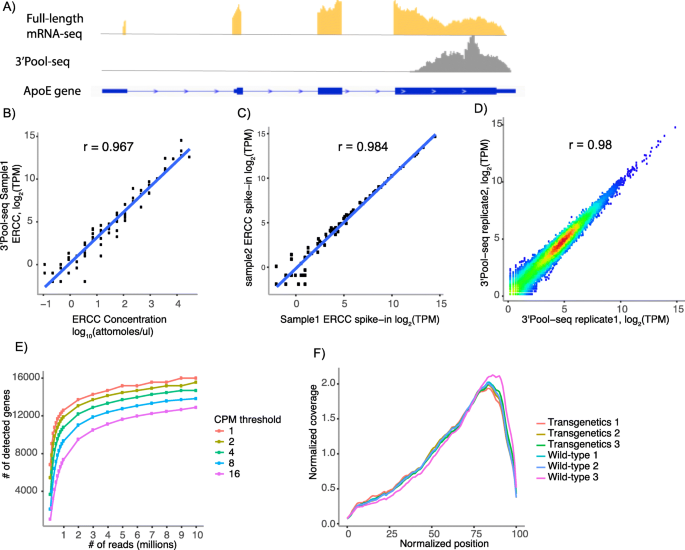
3'Pool-seq: an optimized cost-efficient and scalable method of whole-transcriptome gene expression profiling | BMC Genomics | Full Text